Chickpea ascochyta - evidence that varieties do differ in susceptibility of pods
Author: Kevin Moore, Kristy Hobson, Steve Harden, Paul Nash, Gail Chiplin and Sean Bithell, Department of Primary Industries, NSW, Tamworth | Date: 25 Feb 2015
Take home message
- The susceptibility of pods to Ascochyta Blight is important as infection can cause pod abortion, blemish or kill seed, infected seed is also an inoculum source
- Field trial results indicate that the varietal resistance of chickpea pods are similar to that of vegetative tissue
Background
Understanding the susceptibility of chickpea pod tissue to Ascochyta Blight (caused by Phoma rabiei formerly known as Ascochyta rabiei) is important. Because if pods get infected early in their development they will abort; if fully developed pods get infected near the peduncle (as many do because the calyx holds water), they will abort; pods with developing seeds will either abort, or the seed becomes infected and is killed or the seed becomes infected, but remains viable and is a potential source of inoculum to initiate an epidemic.
Current Australian chickpea varieties and advanced breeding lines differ in susceptibility of their vegetative plant tissues to Ascochyta Blight (see paper on VMP14 trial in these proceedings). However, the chickpea industry believes that pods of all varieties are equally susceptible to Ascochyta (see GRDC's Chickpea disease management factsheet). The 2011 Tamworth chickpea Ascochyta management trial, VMP11 suggested that may not be the case - anecdotal evidence indicated varieties with higher levels of resistance to Ascochyta eg Genesis™ 425 had less disease on their pods. For the past three seasons we have conducted trials at Tamworth designed specifically to capture data on susceptibility of pods of different chickpea genotypes to Ascochyta. Each season we protected the plants with fungicides until 50% podding, then waited for a rain event to inoculate the trials but no rain came. However, the 2014 Tamworth chickpea Ascochyta yield loss trial, VMP14, which was inoculated before flowering, provided an opportunity to collect data on susceptibility of pods of ten genotypes consisting of released varieties and advanced breeding lines.
Methods
Details of VMP14, including disease ratings of the varieties and breeding lines, are reported elsewhere in these proceedings.The trial was inoculated on 15 Jul and re-inoculated on 16 Aug using a new isolate collected from HatTrick at Yallaroi on 24 July. By the end of August, Ascochyta was well established throughout the trial, especially in the unprotected Nil plots (no fungicides). Podding commenced in the 2nd week of September. Eight mm rain fell on 24 Sep followed by 10mm on 25 Sep; 16.4mm fell on 13 Oct with 0.6mm on 14 Oct. On 29 October, 5-6 plants were collected from the outer 2 rows on each side of the 4m wide x 10m long plots. The pods were stripped from each plant, discarding the youngest two pods on each branch (these formed after the last rain event and could not have infected by Ascochyta). The pods were sorted into four classes based on their Ascochyta status: Clean = no Ascochyta lesions; 1 lesion = pods with a single lesion; 2-5 lesions and >5 lesions. A lesion was not called Ascochyta unless pycnidia could be seen either with the naked eye or under a low power dissecting microscope. For each variety the number of pods falling into each of the 4 Aschochyta classes was analysed using ordinal regression. The model estimates (+/- SE) the 3 cut points between the 4 classes and gives a coefficient for each variety.
Results
We acknowledge that this Ascochyta pod data could be confounded, as the plots (JIM and KYB) with the most infected pods and the greater number of lesions were also those that had the highest levels of Ascochyta infection in the vegetative stage. However, we are confident there was sufficient inoculum pressure in the trial. In particular during the two rain events, all pods in the trial would have been exposed to the same aerosol of conidia (40 unsprayed Nil plots in the trial with a combined area of 1600 m2 and an estimated 48,000 infected plants, all with leaf and stem lesions bearing pycnidia). The 2015 trial will hopefully clarify the potential issue of susceptible variety effects on pod infection.
There were large differences in pod infection among the genotypes. Only about 0.3 (30%) of Jimbour and Kyabra pods were clean (no disease), whereas about 96-99% of CICA1007, CICA0912, and GenesisTM 425 pods had no Ascochyta (Table 1). Not only did Jimbour and Kyabra have a greater proportion of Ascochyta infected pods, but these pods were more severely diseased with most of the infected pods having 2-5 or more than 5 Ascochyta lesions (Table 1).
Analysis showed that the varieties can be put into 4 groups with no differences between varieties within a group but significant differences between varieties in different groups. The four groups from least to most susceptible were (C1007,C0912,G425), (BOU,HAT,KAL,MON), (C1211 ) and (JIM,KYB) (Fig. 1).
Table 1. Percentages of pods in each of four Ascochyta categories for ten genotypes
| Genotype | %Clean | %1 Lesion | %2-5 Lesions | %>5 Lesions |
| C0912 | 98.5 | 1.0 | 0.3 | 0.3 |
| C1007 | 97.2 | 1.5 | 1.0 | 0.3 |
| G425 | 96.8 | 2.5 | 0.3 | 0.5 |
| KAL | 86.7 | 7.5 | 5.5 | 0.3 |
| HAT | 86.2 | 9.3 | 4.0 | 0.5 |
| MON | 86.2 | 7.8 | 3.3 | 2.8 |
| BOU | 84.3 | 5.5 | 6.3 | 4.0 |
| C1211 | 67.2 | 13.8 | 14.5 | 4.5 |
| KYB | 33.8 | 15.5 | 30.5 | 20.3 |
| JIM | 28.6 | 21.8 | 31.3 | 18.4 |
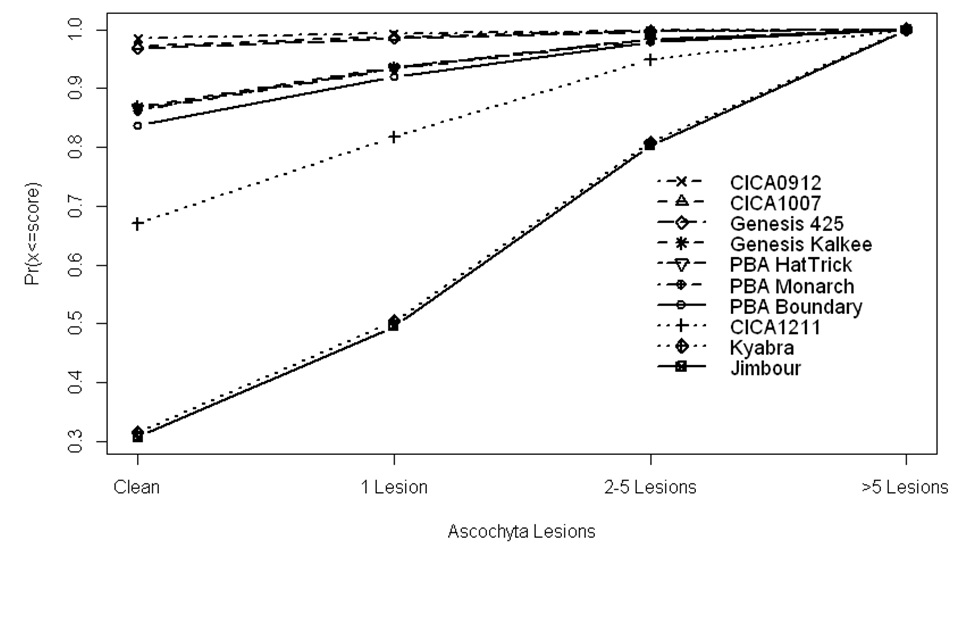
Figure 1. Predicted cumulative proportions of pods for each of four categories of Ascochyta lesions for ten chickpea genotypes in VMP14 trial
Key pod infection findings of VMP14 were:
- Genotypes differed in the number of Ascochyta infected pods.
- Genotypes differed in the severity of Ascochyta on infected pods.
- The ten genotypes fell into four significantly distinct groups in the four pod disease categories with (C1007, C0912, G425) > (BOU, HAT, KAL, MON) > C1211 > (JIM, KYB)
- This grouping agrees closely with current Ascochyta ratings for vegetative tissues
Further information
Further information on chickpea disease management can be found at the Pulse Australia website www.pulseaus.com.au and in the NSW DPI 2015 Winter Crop Variety Sowing Guide eg:
- Chickpea Ascochyta Blight Management pdf
- 2011 Chickpea Disease Management Considerations pdf
- Chickpea Botrytis Grey Mould Management pdf
- Chickpea Integrated Disease Management pdf
- Chickpea Phytophthora Root Rot Management pdf
- Virus Control in Chickpea pdf
- Pulse Seed treatments & Foliar Fungicides pdf
Acknowledgements
This research is made possible by the significant contributions of growers through both trial cooperation, field access and the support of the GRDC; the authors most gratefully thank them and the GRDC. We also thank agronomists for help with the crop inspections and submitting specimens, Dr Mal Ryley, USQ for scientific discussion and advice, Gordon Cumming, Pulse Australia for industry liaison and chemical companies who provide products for research purposes and trial management.
Contact details
Kevin Moore
Department of Primary Industries, Tamworth, NSW
Ph: 02 6763 1133
Mb: 0488 251 866
Fx: 02 6763 1100
Email: kevin.moore@dpi.nsw.gov.au
Was this page helpful?
YOUR FEEDBACK
